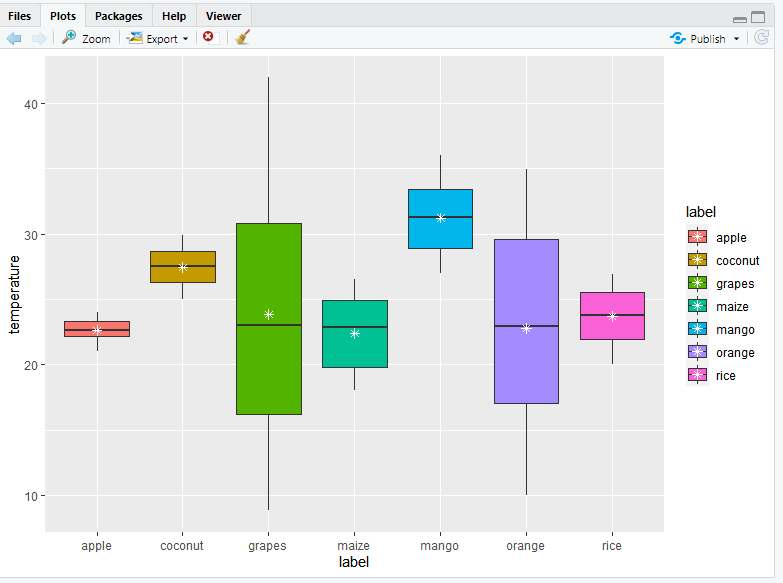
An important analysis question is the quantification and statistical inference of systematic changes between conditions, as compared to within-condition variability. Analogous data also arise for other assay types, including comparative ChIP-Seq, HiC, shRNA screening, and mass spectrometry. The count data are presented as a table which reports, for each sample, the number of sequence fragments that have been assigned to each gene. To do that, we start with aesthetics x=continent, and y=lifeExp and make a boxplot with jittered data points, then we add facet_wrap as layer with year as its argument.A basic task in the analysis of count data from RNA-seq is the detection of differentially expressed genes. Let us make a grouped boxplot such that we have boxplots of lifeExp vs continent for every year. In our case, we can use the function facet_wrap to make grouped boxplots. facet-ing functons in ggplot2 offers general solution to split up the data by one or more variables and make plots with subsets of data together. Geom_point(position=position_jitterdodge(),alpha=0.3) +Ĭustomizing Grouped Boxplot in R Grouped Boxplots with facets in ggplot2Īnother way to make grouped boxplot is to use facet in ggplot. Ggplot(aes(x=continent,y=lifeExp, fill=factor(year))) + So, we use geom_point(position=position_jitterdodge()) here. Similarly, if we use geom_jitter(), the width of the jitter is bit hard to adjust. If we simply add geom_point() as a layer, it will the original data points, but not for every grouped boxplots. Like before, we will jitter the data points. Second, let us show actual data points on the boxplot. We can change the legend to what we want by using the layer labs with fill argument, labs(fill = “Year”) Let us fix that by replacing it to just Year. Let us do three simple customizations.įirst, note that legend of the grouped boxplot we just made still says “factor(year)”. Let us customize the grouped boxplot a bit. Grouped Boxplot in ggplot2 Customizing Grouped Boxplot with ggplot2 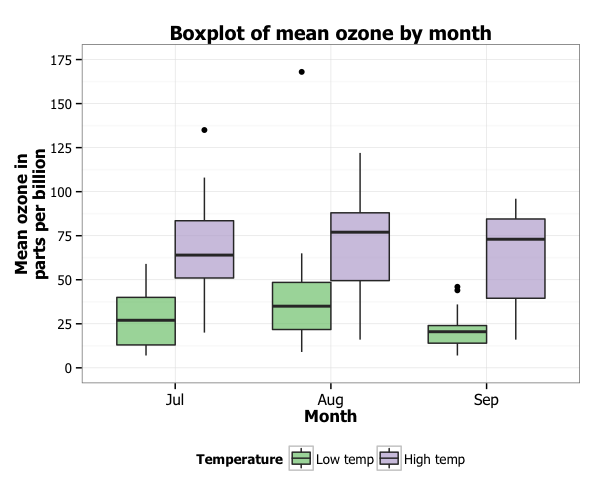
For each continent, we have three boxplots one for each year. Now we have a nice grouped boxplot as we originally intended. Ggplot(aes(x=continent, y=lifeExp, fill=factor(year))) + We can specify that the year is categorical variable by using factor(year) and giving that to the fill argument inside aesthetics. In order to make grouped boxplot using ggplot2, the group variable should be a categorical variable not numerical. Grouped Boxplot: First Try Making Grouped Boxplot with ggplot2 That is the reason we did not get the grouped boxplot. The reason is that if you look at the type of the variable “year” (see with head(gapminder)), you can see that the variable year is of “int” type. However, the resulting boxplot is just a simple boxplot, not a grouped boxplot as we wanted. Ggplot(aes(x=continent, y=lifeExp, fill=year)) + For the sake of simplicity, we just have one geom layer geom_boxplot(). Since we want to use year as grouping variable, we can simply specify “fill=year” in addition to the x-axis and y-axis, and make a boxplot with geom_boxplot(), as shown below. We will use filter function in dplyr to filter the data for the three years of interest and feed the resulting data frame for making a grouped boxplot. 
Let us make a simpler data frame with just data for three years, 1952,1987, and 2007. 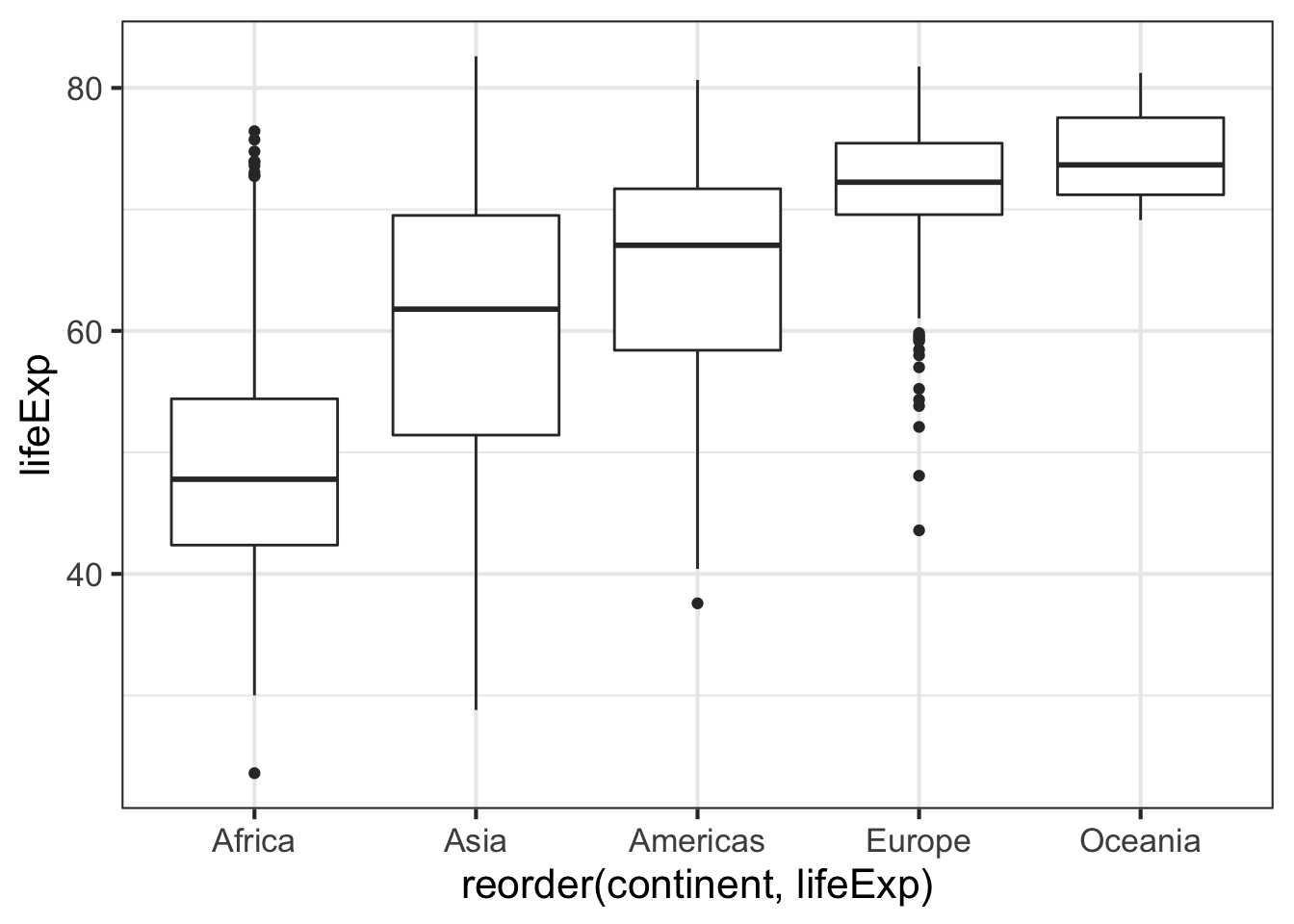
Our gapminder data frame has year variable and has data from multiple years. Let us say, we want to make a grouped boxplot showing the life expectancy over multiple years for each continent.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
